Test ESAWorldCover Classification API#
import requests
from tqdm import tqdm
import geopandas as gpd
import numpy as np
import pandas as pd
import pystac_client
import stackstac
import torch
from rasterio.enums import Resampling
from shapely import Point
import torch
import torch.nn.functional as F
from scipy.ndimage import zoom
import torch.nn as nn
import matplotlib.pyplot as plt
from matplotlib.colors import ListedColormap, BoundaryNorm
import rasterio
lat, lon = 38.25, 21.25
start = "2022-01-01"
end = "2022-01-31"
STAC_API = "https://earth-search.aws.element84.com/v1"
COLLECTION = "sentinel-2-l2a"
# Search the catalogue
catalog = pystac_client.Client.open(STAC_API)
search = catalog.search(
collections=[COLLECTION],
datetime=f"{start}/{end}",
bbox=(lon - 1e-3, lat - 1e-3, lon + 1e-3, lat + 1e-3),
max_items=100,
query={
"eo:cloud_cover": {"lt": 20},
"s2:nodata_pixel_percentage": {"lt": 10},
},
)
all_items = search.get_all_items()
items = []
granules = []
for item in all_items:
if item.properties["s2:granule_id"] not in granules:
items.append(item)
granules.append(item.properties["s2:granule_id"])
print("Found %d Sentinel-2-L2A items" % len(items))
/Users/syam/virtualenvs/myvenv/lib/python3.13/site-packages/pystac_client/item_search.py:896: FutureWarning: get_all_items() is deprecated, use item_collection() instead.
warnings.warn(
Found 4 Sentinel-2-L2A items
proj = items[0].properties["proj:code"]
poidf = gpd.GeoDataFrame(
pd.DataFrame(),
crs="EPSG:4326",
geometry=[Point(lon, lat)],
).to_crs(proj)
coords = poidf.iloc[0].geometry.coords[0]
# Create bounds in projection
size = 2047
gsd = 20
bounds = (
coords[0] - (size * gsd) // 2,
coords[1] - (size * gsd) // 2,
coords[0] + (size * gsd) // 2,
coords[1] + (size * gsd) // 2,
)
assets = ["blue", "green", "red", "nir", "swir16", "scl"]
ds = stackstac.stack(
items,
assets=assets,
bounds=bounds, # pyright: ignore
resolution=gsd,
epsg=int(proj.split(":")[-1]),
dtype="float64", # pyright: ignore
rescale=False,
snap_bounds=True,
resampling=Resampling.nearest,
chunksize=(1, 1, 512, 512),
)
cloud_values = [0, 1, 2, 3, 8, 9, 10] # cloud, shadows, cirrus, etc.
scl_mask = ds.sel(band="scl")
bands = ds.sel(band=["blue", "green", "red", "nir", "swir16"])
valid_mask = ~scl_mask.isin(cloud_values)
bands_masked = bands.where(valid_mask)
median_ds = bands_masked.groupby("time.month").median("time", skipna=True)
median_ds = median_ds.fillna(0)
median_ds = median_ds.compute()
/Users/syam/virtualenvs/myvenv/lib/python3.13/site-packages/dask/_task_spec.py:744: RuntimeWarning: All-NaN slice encountered
return self.func(*new_argspec, **kwargs)
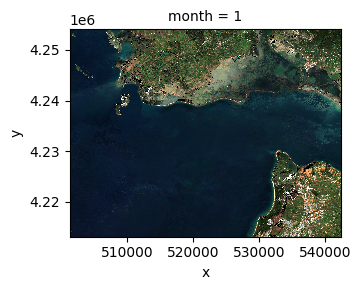
median_ds.sel(band=["red", "green", "blue"]).plot.imshow(
row="month", rgb="band", vmin=0, vmax=2000, col_wrap=6
)
<xarray.plot.facetgrid.FacetGrid at 0x167130ad0>
pixels = torch.from_numpy(median_ds.data.astype(np.float32))
pixels.shape
torch.Size([1, 5, 2048, 2048])
patch_size = 64
stride = 64
batch_size, bands, height, width = pixels.shape
patches = F.unfold(
pixels, kernel_size=patch_size, stride=stride
) # (BATCH, BANDS*PATCH_SIZE*PATCH_SIZE, NUM_PATCHES)
patches = patches.permute(0, 2, 1) # (BATCH, NUM_PATCHES, BANDS*PATCH_SIZE*PATCH_SIZE)
patches = patches.view(
batch_size, -1, bands, patch_size, patch_size
) # (BATCH, NUM_PATCHES, BANDS, PATCH_SIZE, PATCH_SIZE)
patches = patches.reshape(-1, 5, 64, 64)
print(patches.shape)
torch.Size([1024, 5, 64, 64])
B, C, H, W = pixels.shape
#url = "http://localhost:8000/predict"
url = "https://molitserve.internal.meditwin-project.eu/predict"
num_patches = patches.shape[0]
patch_preds = []
for i in tqdm(range(num_patches)):
patch = patches[i,...].numpy() # shape (C, H, W)
patch_list = patch.tolist()
response = requests.post(url, json={"image": patch_list})
if response.status_code != 200:
print(f"Failed at patch {i}: {response.text}")
continue
predicted_mask = response.json()["output"] # shape (H, W)
patch_preds.append(np.array(predicted_mask)) # flatten
100%|██████████| 1024/1024 [07:31<00:00, 2.27it/s]
patch_preds = np.array(patch_preds)
num_patches_per_img = (H // patch_size) * (W // patch_size)
masks = torch.from_numpy(patch_preds).to(torch.float32)
masks = masks.reshape(B, num_patches_per_img, patch_size, patch_size)
masks = masks.reshape(B, num_patches_per_img, -1).permute(0, 2, 1)
reconstructed = F.fold(masks, output_size=(H, W), kernel_size=patch_size, stride=stride)
reconstructed = reconstructed.squeeze(1)
reconstructed = reconstructed.cpu().numpy()[0, ...]
cmap = ListedColormap(
[
"black", # 0 = ignore
"yellow", # 1 = other
"blue", # 2 = water
"red", # 3 = building
]
)
bounds = np.arange(0, 5)
norm = BoundaryNorm(bounds, cmap.N)
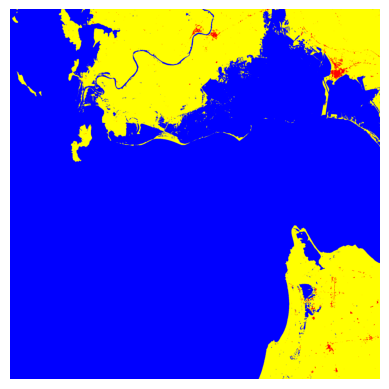
plt.imshow(reconstructed, cmap=cmap, norm=norm)
plt.axis("off")
plt.show()