ESA WorldCover Classification Inference#
In this notebook, we will use the model trained using Sentinel-2 L2A data and ESA WorldCover from 2020 to perform inference on Sentinel-2 images from 2022. The model will classify the land cover types in the images based on the classes specified during the model training.
We will use a combination of what we have learned so far during this training STAC catalogs, StackSTAC, xarray, and PyTorch.
Import libraries#
import geopandas as gpd
import numpy as np
import pandas as pd
import pystac_client
import stackstac
import torch
from rasterio.enums import Resampling
from shapely import Point
import torch
import torch.nn.functional as F
from scipy.ndimage import zoom
import torch.nn as nn
import matplotlib.pyplot as plt
from matplotlib.colors import ListedColormap, BoundaryNorm
import rasterio
Setup the model architecture (the same as training)#
class UNet(nn.Module):
def __init__(self, in_channels=4, out_classes=11, dropout=0.1, init_weights=True):
super(UNet, self).__init__()
def conv_block(in_ch, out_ch):
return nn.Sequential(
nn.Conv2d(in_ch, out_ch, 3, padding=1),
nn.BatchNorm2d(out_ch),
nn.ReLU(inplace=True),
nn.Conv2d(out_ch, out_ch, 3, padding=1),
nn.BatchNorm2d(out_ch),
nn.ReLU(inplace=True),
nn.Dropout2d(dropout),
)
# Encoder
self.enc1 = conv_block(in_channels, 64)
self.enc2 = conv_block(64, 128)
self.enc3 = conv_block(128, 256)
self.pool = nn.MaxPool2d(2)
# Bottleneck
self.bottleneck = conv_block(256, 512)
# Decoder
self.dec3 = conv_block(512 + 256, 256)
self.dec2 = conv_block(256 + 128, 128)
self.dec1 = conv_block(128 + 64, 64)
self.dec0 = conv_block(64, 32)
# Final classifier
self.final = nn.Conv2d(32, out_classes, kernel_size=1)
if init_weights:
self._init_weights()
def _init_weights(self):
for m in self.modules():
if isinstance(m, nn.Conv2d):
nn.init.kaiming_normal_(m.weight, nonlinearity="relu")
if m.bias is not None:
nn.init.zeros_(m.bias)
def forward(self, x):
e1 = self.enc1(x) # (B, 64, H, W)
e2 = self.enc2(self.pool(e1)) # (B, 128, H/2, W/2)
e3 = self.enc3(self.pool(e2)) # (B, 256, H/4, W/4)
b = self.bottleneck(self.pool(e3)) # (B, 512, H/8, W/8)
d3 = self._upsample_concat(b, e3) # (B, 512+256, H/4, W/4)
d3 = self.dec3(d3)
d2 = self._upsample_concat(d3, e2) # (B, 256+128, H/2, W/2)
d2 = self.dec2(d2)
d1 = self._upsample_concat(d2, e1) # (B, 128+64, H, W)
d1 = self.dec1(d1)
d0 = self.dec0(d1) # (B, 32, H, W)
return self.final(d0) # logits: (B, out_classes, H, W)
def _upsample_concat(self, x, skip):
x = F.interpolate(x, size=skip.shape[2:], mode="bilinear", align_corners=False)
return torch.cat([x, skip], dim=1)
Search STAC catalog for Sentinel-2 L2A images#
lat, lon = 38.25, 21.25
start = "2022-01-01"
end = "2022-01-31"
STAC_API = "https://earth-search.aws.element84.com/v1"
COLLECTION = "sentinel-2-l2a"
# Search the catalogue
catalog = pystac_client.Client.open(STAC_API)
search = catalog.search(
collections=[COLLECTION],
datetime=f"{start}/{end}",
bbox=(lon - 1e-3, lat - 1e-3, lon + 1e-3, lat + 1e-3),
max_items=100,
query={
"eo:cloud_cover": {"lt": 20},
"s2:nodata_pixel_percentage": {"lt": 10},
},
)
all_items = search.get_all_items()
items = []
granules = []
for item in all_items:
if item.properties["s2:granule_id"] not in granules:
items.append(item)
granules.append(item.properties["s2:granule_id"])
print("Found %d Sentinel-2-L2A items" %len(items))
/Users/syam/virtualenvs/myvenv/lib/python3.13/site-packages/pystac_client/item_search.py:896: FutureWarning: get_all_items() is deprecated, use item_collection() instead.
warnings.warn(
Found 4 Sentinel-2-L2A items
items[0]
- type "Feature"
- stac_version "1.1.0"
stac_extensions[] 7 items
- 0 "https://stac-extensions.github.io/view/v1.0.0/schema.json"
- 1 "https://stac-extensions.github.io/mgrs/v1.0.0/schema.json"
- 2 "https://stac-extensions.github.io/grid/v1.1.0/schema.json"
- 3 "https://stac-extensions.github.io/raster/v1.1.0/schema.json"
- 4 "https://stac-extensions.github.io/processing/v1.1.0/schema.json"
- 5 "https://stac-extensions.github.io/projection/v2.0.0/schema.json"
- 6 "https://stac-extensions.github.io/eo/v1.1.0/schema.json"
- id "S2A_34SEH_20220123_0_L2A"
geometry
- type "Polygon"
coordinates[] 1 items
0[] 5 items
0[] 2 items
- 0 20.99978104422428
- 1 38.84898947181679
1[] 2 items
- 0 22.264948435187954
- 1 38.842139787815405
2[] 2 items
- 0 22.24785419613706
- 1 37.852838634285284
3[] 2 items
- 0 20.99978400408747
- 1 37.85945097203489
4[] 2 items
- 0 20.99978104422428
- 1 38.84898947181679
bbox[] 4 items
- 0 20.99978104422428
- 1 37.852838634285284
- 2 22.264948435187954
- 3 38.84898947181679
properties
- created "2022-11-06T12:05:37.418Z"
- platform "sentinel-2a"
- constellation "sentinel-2"
instruments[] 1 items
- 0 "msi"
- eo:cloud_cover 5.098246
- mgrs:utm_zone 34
- mgrs:latitude_band "S"
- mgrs:grid_square "EH"
- grid:code "MGRS-34SEH"
- view:sun_azimuth 159.499336052921
- view:sun_elevation 29.593954613099
- s2:degraded_msi_data_percentage 0
- s2:nodata_pixel_percentage 8.6e-05
- s2:saturated_defective_pixel_percentage 0
- s2:dark_features_percentage 6.747229
- s2:cloud_shadow_percentage 2.256003
- s2:vegetation_percentage 47.629133
- s2:not_vegetated_percentage 5.207532
- s2:water_percentage 24.151275
- s2:unclassified_percentage 5.07446
- s2:medium_proba_clouds_percentage 1.950639
- s2:high_proba_clouds_percentage 1.440859
- s2:thin_cirrus_percentage 1.706748
- s2:snow_ice_percentage 3.836122
- s2:product_type "S2MSI2A"
- s2:processing_baseline "03.01"
- s2:product_uri "S2A_MSIL2A_20220123T092301_N0301_R093_T34SEH_20220123T122825.SAFE"
- s2:generation_time "2022-01-23T12:28:25.000000Z"
- s2:datatake_id "GS2A_20220123T092301_034410_N03.01"
- s2:datatake_type "INS-NOBS"
- s2:datastrip_id "S2A_OPER_MSI_L2A_DS_VGS2_20220123T122825_S20220123T092339_N03.01"
- s2:granule_id "S2A_OPER_MSI_L2A_TL_VGS2_20220123T122825_A034410_T34SEH_N03.01"
- s2:reflectance_conversion_factor 1.03296508738509
- datetime "2022-01-23T09:29:54.217000Z"
- s2:sequence "0"
- earthsearch:s3_path "s3://sentinel-cogs/sentinel-s2-l2a-cogs/34/S/EH/2022/1/S2A_34SEH_20220123_0_L2A"
- earthsearch:payload_id "roda-sentinel2/workflow-sentinel2-to-stac/07840e2fd6554cd332b3f1f43ab2591f"
- earthsearch:boa_offset_applied False
processing:software
- sentinel2-to-stac "0.1.0"
- updated "2022-11-06T12:05:37.418Z"
- proj:code "EPSG:32634"
links[] 8 items
0
- rel "self"
- href "https://earth-search.aws.element84.com/v1/collections/sentinel-2-l2a/items/S2A_34SEH_20220123_0_L2A"
- type "application/geo+json"
1
- rel "canonical"
- href "s3://sentinel-cogs/sentinel-s2-l2a-cogs/34/S/EH/2022/1/S2A_34SEH_20220123_0_L2A/S2A_34SEH_20220123_0_L2A.json"
- type "application/json"
2
- rel "license"
- href "https://sentinel.esa.int/documents/247904/690755/Sentinel_Data_Legal_Notice"
3
- rel "derived_from"
- href "https://earth-search.aws.element84.com/v1/collections/sentinel-2-l1c/items/S2A_34SEH_20220123_0_L1C"
- type "application/geo+json"
4
- rel "parent"
- href "https://earth-search.aws.element84.com/v1/collections/sentinel-2-l2a"
- type "application/json"
5
- rel "collection"
- href "https://earth-search.aws.element84.com/v1/collections/sentinel-2-l2a"
- type "application/json"
6
- rel "root"
- href "https://earth-search.aws.element84.com/v1"
- type "application/json"
- title "Earth Search by Element 84"
7
- rel "thumbnail"
- href "https://earth-search.aws.element84.com/v1/collections/sentinel-2-l2a/items/S2A_34SEH_20220123_0_L2A/thumbnail"
assets
aot
- href "https://sentinel-cogs.s3.us-west-2.amazonaws.com/sentinel-s2-l2a-cogs/34/S/EH/2022/1/S2A_34SEH_20220123_0_L2A/AOT.tif"
- type "image/tiff; application=geotiff; profile=cloud-optimized"
- title "Aerosol optical thickness (AOT)"
proj:shape[] 2 items
- 0 5490
- 1 5490
proj:transform[] 6 items
- 0 20
- 1 0
- 2 499980
- 3 0
- 4 -20
- 5 4300020
raster:bands[] 1 items
0
- nodata 0
- data_type "uint16"
- bits_per_sample 15
- spatial_resolution 20
- scale 0.001
- offset 0
roles[] 2 items
- 0 "data"
- 1 "reflectance"
blue
- href "https://sentinel-cogs.s3.us-west-2.amazonaws.com/sentinel-s2-l2a-cogs/34/S/EH/2022/1/S2A_34SEH_20220123_0_L2A/B02.tif"
- type "image/tiff; application=geotiff; profile=cloud-optimized"
- title "Blue (band 2) - 10m"
eo:bands[] 1 items
0
- name "blue"
- common_name "blue"
- description "Blue (band 2)"
- center_wavelength 0.49
- full_width_half_max 0.098
- gsd 10
proj:shape[] 2 items
- 0 10980
- 1 10980
proj:transform[] 6 items
- 0 10
- 1 0
- 2 499980
- 3 0
- 4 -10
- 5 4300020
raster:bands[] 1 items
0
- nodata 0
- data_type "uint16"
- bits_per_sample 15
- spatial_resolution 10
- scale 0.0001
- offset 0
roles[] 2 items
- 0 "data"
- 1 "reflectance"
coastal
- href "https://sentinel-cogs.s3.us-west-2.amazonaws.com/sentinel-s2-l2a-cogs/34/S/EH/2022/1/S2A_34SEH_20220123_0_L2A/B01.tif"
- type "image/tiff; application=geotiff; profile=cloud-optimized"
- title "Coastal aerosol (band 1) - 60m"
eo:bands[] 1 items
0
- name "coastal"
- common_name "coastal"
- description "Coastal aerosol (band 1)"
- center_wavelength 0.443
- full_width_half_max 0.027
- gsd 60
proj:shape[] 2 items
- 0 1830
- 1 1830
proj:transform[] 6 items
- 0 60
- 1 0
- 2 499980
- 3 0
- 4 -60
- 5 4300020
raster:bands[] 1 items
0
- nodata 0
- data_type "uint16"
- bits_per_sample 15
- spatial_resolution 60
- scale 0.0001
- offset 0
roles[] 2 items
- 0 "data"
- 1 "reflectance"
granule_metadata
- href "https://sentinel-cogs.s3.us-west-2.amazonaws.com/sentinel-s2-l2a-cogs/34/S/EH/2022/1/S2A_34SEH_20220123_0_L2A/granule_metadata.xml"
- type "application/xml"
roles[] 1 items
- 0 "metadata"
green
- href "https://sentinel-cogs.s3.us-west-2.amazonaws.com/sentinel-s2-l2a-cogs/34/S/EH/2022/1/S2A_34SEH_20220123_0_L2A/B03.tif"
- type "image/tiff; application=geotiff; profile=cloud-optimized"
- title "Green (band 3) - 10m"
eo:bands[] 1 items
0
- name "green"
- common_name "green"
- description "Green (band 3)"
- center_wavelength 0.56
- full_width_half_max 0.045
- gsd 10
proj:shape[] 2 items
- 0 10980
- 1 10980
proj:transform[] 6 items
- 0 10
- 1 0
- 2 499980
- 3 0
- 4 -10
- 5 4300020
raster:bands[] 1 items
0
- nodata 0
- data_type "uint16"
- bits_per_sample 15
- spatial_resolution 10
- scale 0.0001
- offset 0
roles[] 2 items
- 0 "data"
- 1 "reflectance"
nir
- href "https://sentinel-cogs.s3.us-west-2.amazonaws.com/sentinel-s2-l2a-cogs/34/S/EH/2022/1/S2A_34SEH_20220123_0_L2A/B08.tif"
- type "image/tiff; application=geotiff; profile=cloud-optimized"
- title "NIR 1 (band 8) - 10m"
eo:bands[] 1 items
0
- name "nir"
- common_name "nir"
- description "NIR 1 (band 8)"
- center_wavelength 0.842
- full_width_half_max 0.145
- gsd 10
proj:shape[] 2 items
- 0 10980
- 1 10980
proj:transform[] 6 items
- 0 10
- 1 0
- 2 499980
- 3 0
- 4 -10
- 5 4300020
raster:bands[] 1 items
0
- nodata 0
- data_type "uint16"
- bits_per_sample 15
- spatial_resolution 10
- scale 0.0001
- offset 0
roles[] 2 items
- 0 "data"
- 1 "reflectance"
nir08
- href "https://sentinel-cogs.s3.us-west-2.amazonaws.com/sentinel-s2-l2a-cogs/34/S/EH/2022/1/S2A_34SEH_20220123_0_L2A/B8A.tif"
- type "image/tiff; application=geotiff; profile=cloud-optimized"
- title "NIR 2 (band 8A) - 20m"
eo:bands[] 1 items
0
- name "nir08"
- common_name "nir08"
- description "NIR 2 (band 8A)"
- center_wavelength 0.865
- full_width_half_max 0.033
- gsd 20
proj:shape[] 2 items
- 0 5490
- 1 5490
proj:transform[] 6 items
- 0 20
- 1 0
- 2 499980
- 3 0
- 4 -20
- 5 4300020
raster:bands[] 1 items
0
- nodata 0
- data_type "uint16"
- bits_per_sample 15
- spatial_resolution 20
- scale 0.0001
- offset 0
roles[] 2 items
- 0 "data"
- 1 "reflectance"
nir09
- href "https://sentinel-cogs.s3.us-west-2.amazonaws.com/sentinel-s2-l2a-cogs/34/S/EH/2022/1/S2A_34SEH_20220123_0_L2A/B09.tif"
- type "image/tiff; application=geotiff; profile=cloud-optimized"
- title "NIR 3 (band 9) - 60m"
eo:bands[] 1 items
0
- name "nir09"
- common_name "nir09"
- description "NIR 3 (band 9)"
- center_wavelength 0.945
- full_width_half_max 0.026
- gsd 60
proj:shape[] 2 items
- 0 1830
- 1 1830
proj:transform[] 6 items
- 0 60
- 1 0
- 2 499980
- 3 0
- 4 -60
- 5 4300020
raster:bands[] 1 items
0
- nodata 0
- data_type "uint16"
- bits_per_sample 15
- spatial_resolution 60
- scale 0.0001
- offset 0
roles[] 2 items
- 0 "data"
- 1 "reflectance"
red
- href "https://sentinel-cogs.s3.us-west-2.amazonaws.com/sentinel-s2-l2a-cogs/34/S/EH/2022/1/S2A_34SEH_20220123_0_L2A/B04.tif"
- type "image/tiff; application=geotiff; profile=cloud-optimized"
- title "Red (band 4) - 10m"
eo:bands[] 1 items
0
- name "red"
- common_name "red"
- description "Red (band 4)"
- center_wavelength 0.665
- full_width_half_max 0.038
- gsd 10
proj:shape[] 2 items
- 0 10980
- 1 10980
proj:transform[] 6 items
- 0 10
- 1 0
- 2 499980
- 3 0
- 4 -10
- 5 4300020
raster:bands[] 1 items
0
- nodata 0
- data_type "uint16"
- bits_per_sample 15
- spatial_resolution 10
- scale 0.0001
- offset 0
roles[] 2 items
- 0 "data"
- 1 "reflectance"
rededge1
- href "https://sentinel-cogs.s3.us-west-2.amazonaws.com/sentinel-s2-l2a-cogs/34/S/EH/2022/1/S2A_34SEH_20220123_0_L2A/B05.tif"
- type "image/tiff; application=geotiff; profile=cloud-optimized"
- title "Red edge 1 (band 5) - 20m"
eo:bands[] 1 items
0
- name "rededge1"
- common_name "rededge"
- description "Red edge 1 (band 5)"
- center_wavelength 0.704
- full_width_half_max 0.019
- gsd 20
proj:shape[] 2 items
- 0 5490
- 1 5490
proj:transform[] 6 items
- 0 20
- 1 0
- 2 499980
- 3 0
- 4 -20
- 5 4300020
raster:bands[] 1 items
0
- nodata 0
- data_type "uint16"
- bits_per_sample 15
- spatial_resolution 20
- scale 0.0001
- offset 0
roles[] 2 items
- 0 "data"
- 1 "reflectance"
rededge2
- href "https://sentinel-cogs.s3.us-west-2.amazonaws.com/sentinel-s2-l2a-cogs/34/S/EH/2022/1/S2A_34SEH_20220123_0_L2A/B06.tif"
- type "image/tiff; application=geotiff; profile=cloud-optimized"
- title "Red edge 2 (band 6) - 20m"
eo:bands[] 1 items
0
- name "rededge2"
- common_name "rededge"
- description "Red edge 2 (band 6)"
- center_wavelength 0.74
- full_width_half_max 0.018
- gsd 20
proj:shape[] 2 items
- 0 5490
- 1 5490
proj:transform[] 6 items
- 0 20
- 1 0
- 2 499980
- 3 0
- 4 -20
- 5 4300020
raster:bands[] 1 items
0
- nodata 0
- data_type "uint16"
- bits_per_sample 15
- spatial_resolution 20
- scale 0.0001
- offset 0
roles[] 2 items
- 0 "data"
- 1 "reflectance"
rededge3
- href "https://sentinel-cogs.s3.us-west-2.amazonaws.com/sentinel-s2-l2a-cogs/34/S/EH/2022/1/S2A_34SEH_20220123_0_L2A/B07.tif"
- type "image/tiff; application=geotiff; profile=cloud-optimized"
- title "Red edge 3 (band 7) - 20m"
eo:bands[] 1 items
0
- name "rededge3"
- common_name "rededge"
- description "Red edge 3 (band 7)"
- center_wavelength 0.783
- full_width_half_max 0.028
- gsd 20
proj:shape[] 2 items
- 0 5490
- 1 5490
proj:transform[] 6 items
- 0 20
- 1 0
- 2 499980
- 3 0
- 4 -20
- 5 4300020
raster:bands[] 1 items
0
- nodata 0
- data_type "uint16"
- bits_per_sample 15
- spatial_resolution 20
- scale 0.0001
- offset 0
roles[] 2 items
- 0 "data"
- 1 "reflectance"
scl
- href "https://sentinel-cogs.s3.us-west-2.amazonaws.com/sentinel-s2-l2a-cogs/34/S/EH/2022/1/S2A_34SEH_20220123_0_L2A/SCL.tif"
- type "image/tiff; application=geotiff; profile=cloud-optimized"
- title "Scene classification map (SCL)"
proj:shape[] 2 items
- 0 5490
- 1 5490
proj:transform[] 6 items
- 0 20
- 1 0
- 2 499980
- 3 0
- 4 -20
- 5 4300020
raster:bands[] 1 items
0
- nodata 0
- data_type "uint8"
- spatial_resolution 20
roles[] 2 items
- 0 "data"
- 1 "reflectance"
swir16
- href "https://sentinel-cogs.s3.us-west-2.amazonaws.com/sentinel-s2-l2a-cogs/34/S/EH/2022/1/S2A_34SEH_20220123_0_L2A/B11.tif"
- type "image/tiff; application=geotiff; profile=cloud-optimized"
- title "SWIR 1 (band 11) - 20m"
eo:bands[] 1 items
0
- name "swir16"
- common_name "swir16"
- description "SWIR 1 (band 11)"
- center_wavelength 1.61
- full_width_half_max 0.143
- gsd 20
proj:shape[] 2 items
- 0 5490
- 1 5490
proj:transform[] 6 items
- 0 20
- 1 0
- 2 499980
- 3 0
- 4 -20
- 5 4300020
raster:bands[] 1 items
0
- nodata 0
- data_type "uint16"
- bits_per_sample 15
- spatial_resolution 20
- scale 0.0001
- offset 0
roles[] 2 items
- 0 "data"
- 1 "reflectance"
swir22
- href "https://sentinel-cogs.s3.us-west-2.amazonaws.com/sentinel-s2-l2a-cogs/34/S/EH/2022/1/S2A_34SEH_20220123_0_L2A/B12.tif"
- type "image/tiff; application=geotiff; profile=cloud-optimized"
- title "SWIR 2 (band 12) - 20m"
eo:bands[] 1 items
0
- name "swir22"
- common_name "swir22"
- description "SWIR 2 (band 12)"
- center_wavelength 2.19
- full_width_half_max 0.242
- gsd 20
proj:shape[] 2 items
- 0 5490
- 1 5490
proj:transform[] 6 items
- 0 20
- 1 0
- 2 499980
- 3 0
- 4 -20
- 5 4300020
raster:bands[] 1 items
0
- nodata 0
- data_type "uint16"
- bits_per_sample 15
- spatial_resolution 20
- scale 0.0001
- offset 0
roles[] 2 items
- 0 "data"
- 1 "reflectance"
thumbnail
- href "https://sentinel-cogs.s3.us-west-2.amazonaws.com/sentinel-s2-l2a-cogs/34/S/EH/2022/1/S2A_34SEH_20220123_0_L2A/thumbnail.jpg"
- type "image/jpeg"
- title "Thumbnail image"
roles[] 1 items
- 0 "thumbnail"
tileinfo_metadata
- href "https://sentinel-cogs.s3.us-west-2.amazonaws.com/sentinel-s2-l2a-cogs/34/S/EH/2022/1/S2A_34SEH_20220123_0_L2A/tileinfo_metadata.json"
- type "application/json"
roles[] 1 items
- 0 "metadata"
visual
- href "https://sentinel-cogs.s3.us-west-2.amazonaws.com/sentinel-s2-l2a-cogs/34/S/EH/2022/1/S2A_34SEH_20220123_0_L2A/TCI.tif"
- type "image/tiff; application=geotiff; profile=cloud-optimized"
- title "True color image"
eo:bands[] 3 items
0
- name "red"
- common_name "red"
- description "Red (band 4)"
- center_wavelength 0.665
- full_width_half_max 0.038
1
- name "green"
- common_name "green"
- description "Green (band 3)"
- center_wavelength 0.56
- full_width_half_max 0.045
2
- name "blue"
- common_name "blue"
- description "Blue (band 2)"
- center_wavelength 0.49
- full_width_half_max 0.098
proj:shape[] 2 items
- 0 10980
- 1 10980
proj:transform[] 6 items
- 0 10
- 1 0
- 2 499980
- 3 0
- 4 -10
- 5 4300020
roles[] 1 items
- 0 "visual"
wvp
- href "https://sentinel-cogs.s3.us-west-2.amazonaws.com/sentinel-s2-l2a-cogs/34/S/EH/2022/1/S2A_34SEH_20220123_0_L2A/WVP.tif"
- type "image/tiff; application=geotiff; profile=cloud-optimized"
- title "Water vapour (WVP)"
proj:shape[] 2 items
- 0 5490
- 1 5490
proj:transform[] 6 items
- 0 20
- 1 0
- 2 499980
- 3 0
- 4 -20
- 5 4300020
raster:bands[] 1 items
0
- nodata 0
- data_type "uint16"
- bits_per_sample 15
- spatial_resolution 20
- unit "cm"
- scale 0.001
- offset 0
roles[] 2 items
- 0 "data"
- 1 "reflectance"
aot-jp2
- href "s3://sentinel-s2-l2a/tiles/34/S/EH/2022/1/23/0/AOT.jp2"
- type "image/jp2"
- title "Aerosol optical thickness (AOT)"
proj:shape[] 2 items
- 0 5490
- 1 5490
proj:transform[] 6 items
- 0 20
- 1 0
- 2 499980
- 3 0
- 4 -20
- 5 4300020
raster:bands[] 1 items
0
- nodata 0
- data_type "uint16"
- bits_per_sample 15
- spatial_resolution 20
- scale 0.001
- offset 0
roles[] 2 items
- 0 "data"
- 1 "reflectance"
blue-jp2
- href "s3://sentinel-s2-l2a/tiles/34/S/EH/2022/1/23/0/B02.jp2"
- type "image/jp2"
- title "Blue (band 2) - 10m"
eo:bands[] 1 items
0
- name "blue"
- common_name "blue"
- description "Blue (band 2)"
- center_wavelength 0.49
- full_width_half_max 0.098
- gsd 10
proj:shape[] 2 items
- 0 10980
- 1 10980
proj:transform[] 6 items
- 0 10
- 1 0
- 2 499980
- 3 0
- 4 -10
- 5 4300020
raster:bands[] 1 items
0
- nodata 0
- data_type "uint16"
- bits_per_sample 15
- spatial_resolution 10
- scale 0.0001
- offset 0
roles[] 2 items
- 0 "data"
- 1 "reflectance"
coastal-jp2
- href "s3://sentinel-s2-l2a/tiles/34/S/EH/2022/1/23/0/B01.jp2"
- type "image/jp2"
- title "Coastal aerosol (band 1) - 60m"
eo:bands[] 1 items
0
- name "coastal"
- common_name "coastal"
- description "Coastal aerosol (band 1)"
- center_wavelength 0.443
- full_width_half_max 0.027
- gsd 60
proj:shape[] 2 items
- 0 1830
- 1 1830
proj:transform[] 6 items
- 0 60
- 1 0
- 2 499980
- 3 0
- 4 -60
- 5 4300020
raster:bands[] 1 items
0
- nodata 0
- data_type "uint16"
- bits_per_sample 15
- spatial_resolution 60
- scale 0.0001
- offset 0
roles[] 2 items
- 0 "data"
- 1 "reflectance"
green-jp2
- href "s3://sentinel-s2-l2a/tiles/34/S/EH/2022/1/23/0/B03.jp2"
- type "image/jp2"
- title "Green (band 3) - 10m"
eo:bands[] 1 items
0
- name "green"
- common_name "green"
- description "Green (band 3)"
- center_wavelength 0.56
- full_width_half_max 0.045
- gsd 10
proj:shape[] 2 items
- 0 10980
- 1 10980
proj:transform[] 6 items
- 0 10
- 1 0
- 2 499980
- 3 0
- 4 -10
- 5 4300020
raster:bands[] 1 items
0
- nodata 0
- data_type "uint16"
- bits_per_sample 15
- spatial_resolution 10
- scale 0.0001
- offset 0
roles[] 2 items
- 0 "data"
- 1 "reflectance"
nir-jp2
- href "s3://sentinel-s2-l2a/tiles/34/S/EH/2022/1/23/0/B08.jp2"
- type "image/jp2"
- title "NIR 1 (band 8) - 10m"
eo:bands[] 1 items
0
- name "nir"
- common_name "nir"
- description "NIR 1 (band 8)"
- center_wavelength 0.842
- full_width_half_max 0.145
- gsd 10
proj:shape[] 2 items
- 0 10980
- 1 10980
proj:transform[] 6 items
- 0 10
- 1 0
- 2 499980
- 3 0
- 4 -10
- 5 4300020
raster:bands[] 1 items
0
- nodata 0
- data_type "uint16"
- bits_per_sample 15
- spatial_resolution 10
- scale 0.0001
- offset 0
roles[] 2 items
- 0 "data"
- 1 "reflectance"
nir08-jp2
- href "s3://sentinel-s2-l2a/tiles/34/S/EH/2022/1/23/0/B8A.jp2"
- type "image/jp2"
- title "NIR 2 (band 8A) - 20m"
eo:bands[] 1 items
0
- name "nir08"
- common_name "nir08"
- description "NIR 2 (band 8A)"
- center_wavelength 0.865
- full_width_half_max 0.033
- gsd 20
proj:shape[] 2 items
- 0 5490
- 1 5490
proj:transform[] 6 items
- 0 20
- 1 0
- 2 499980
- 3 0
- 4 -20
- 5 4300020
raster:bands[] 1 items
0
- nodata 0
- data_type "uint16"
- bits_per_sample 15
- spatial_resolution 20
- scale 0.0001
- offset 0
roles[] 2 items
- 0 "data"
- 1 "reflectance"
nir09-jp2
- href "s3://sentinel-s2-l2a/tiles/34/S/EH/2022/1/23/0/B09.jp2"
- type "image/jp2"
- title "NIR 3 (band 9) - 60m"
eo:bands[] 1 items
0
- name "nir09"
- common_name "nir09"
- description "NIR 3 (band 9)"
- center_wavelength 0.945
- full_width_half_max 0.026
- gsd 60
proj:shape[] 2 items
- 0 1830
- 1 1830
proj:transform[] 6 items
- 0 60
- 1 0
- 2 499980
- 3 0
- 4 -60
- 5 4300020
raster:bands[] 1 items
0
- nodata 0
- data_type "uint16"
- bits_per_sample 15
- spatial_resolution 60
- scale 0.0001
- offset 0
roles[] 2 items
- 0 "data"
- 1 "reflectance"
red-jp2
- href "s3://sentinel-s2-l2a/tiles/34/S/EH/2022/1/23/0/B04.jp2"
- type "image/jp2"
- title "Red (band 4) - 10m"
eo:bands[] 1 items
0
- name "red"
- common_name "red"
- description "Red (band 4)"
- center_wavelength 0.665
- full_width_half_max 0.038
- gsd 10
proj:shape[] 2 items
- 0 10980
- 1 10980
proj:transform[] 6 items
- 0 10
- 1 0
- 2 499980
- 3 0
- 4 -10
- 5 4300020
raster:bands[] 1 items
0
- nodata 0
- data_type "uint16"
- bits_per_sample 15
- spatial_resolution 10
- scale 0.0001
- offset 0
roles[] 2 items
- 0 "data"
- 1 "reflectance"
rededge1-jp2
- href "s3://sentinel-s2-l2a/tiles/34/S/EH/2022/1/23/0/B05.jp2"
- type "image/jp2"
- title "Red edge 1 (band 5) - 20m"
eo:bands[] 1 items
0
- name "rededge1"
- common_name "rededge"
- description "Red edge 1 (band 5)"
- center_wavelength 0.704
- full_width_half_max 0.019
- gsd 20
proj:shape[] 2 items
- 0 5490
- 1 5490
proj:transform[] 6 items
- 0 20
- 1 0
- 2 499980
- 3 0
- 4 -20
- 5 4300020
raster:bands[] 1 items
0
- nodata 0
- data_type "uint16"
- bits_per_sample 15
- spatial_resolution 20
- scale 0.0001
- offset 0
roles[] 2 items
- 0 "data"
- 1 "reflectance"
rededge2-jp2
- href "s3://sentinel-s2-l2a/tiles/34/S/EH/2022/1/23/0/B06.jp2"
- type "image/jp2"
- title "Red edge 2 (band 6) - 20m"
eo:bands[] 1 items
0
- name "rededge2"
- common_name "rededge"
- description "Red edge 2 (band 6)"
- center_wavelength 0.74
- full_width_half_max 0.018
- gsd 20
proj:shape[] 2 items
- 0 5490
- 1 5490
proj:transform[] 6 items
- 0 20
- 1 0
- 2 499980
- 3 0
- 4 -20
- 5 4300020
raster:bands[] 1 items
0
- nodata 0
- data_type "uint16"
- bits_per_sample 15
- spatial_resolution 20
- scale 0.0001
- offset 0
roles[] 2 items
- 0 "data"
- 1 "reflectance"
rededge3-jp2
- href "s3://sentinel-s2-l2a/tiles/34/S/EH/2022/1/23/0/B07.jp2"
- type "image/jp2"
- title "Red edge 3 (band 7) - 20m"
eo:bands[] 1 items
0
- name "rededge3"
- common_name "rededge"
- description "Red edge 3 (band 7)"
- center_wavelength 0.783
- full_width_half_max 0.028
- gsd 20
proj:shape[] 2 items
- 0 5490
- 1 5490
proj:transform[] 6 items
- 0 20
- 1 0
- 2 499980
- 3 0
- 4 -20
- 5 4300020
raster:bands[] 1 items
0
- nodata 0
- data_type "uint16"
- bits_per_sample 15
- spatial_resolution 20
- scale 0.0001
- offset 0
roles[] 2 items
- 0 "data"
- 1 "reflectance"
scl-jp2
- href "s3://sentinel-s2-l2a/tiles/34/S/EH/2022/1/23/0/SCL.jp2"
- type "image/jp2"
- title "Scene classification map (SCL)"
proj:shape[] 2 items
- 0 5490
- 1 5490
proj:transform[] 6 items
- 0 20
- 1 0
- 2 499980
- 3 0
- 4 -20
- 5 4300020
raster:bands[] 1 items
0
- nodata 0
- data_type "uint8"
- spatial_resolution 20
roles[] 2 items
- 0 "data"
- 1 "reflectance"
swir16-jp2
- href "s3://sentinel-s2-l2a/tiles/34/S/EH/2022/1/23/0/B11.jp2"
- type "image/jp2"
- title "SWIR 1 (band 11) - 20m"
eo:bands[] 1 items
0
- name "swir16"
- common_name "swir16"
- description "SWIR 1 (band 11)"
- center_wavelength 1.61
- full_width_half_max 0.143
- gsd 20
proj:shape[] 2 items
- 0 5490
- 1 5490
proj:transform[] 6 items
- 0 20
- 1 0
- 2 499980
- 3 0
- 4 -20
- 5 4300020
raster:bands[] 1 items
0
- nodata 0
- data_type "uint16"
- bits_per_sample 15
- spatial_resolution 20
- scale 0.0001
- offset 0
roles[] 2 items
- 0 "data"
- 1 "reflectance"
swir22-jp2
- href "s3://sentinel-s2-l2a/tiles/34/S/EH/2022/1/23/0/B12.jp2"
- type "image/jp2"
- title "SWIR 2 (band 12) - 20m"
eo:bands[] 1 items
0
- name "swir22"
- common_name "swir22"
- description "SWIR 2 (band 12)"
- center_wavelength 2.19
- full_width_half_max 0.242
- gsd 20
proj:shape[] 2 items
- 0 5490
- 1 5490
proj:transform[] 6 items
- 0 20
- 1 0
- 2 499980
- 3 0
- 4 -20
- 5 4300020
raster:bands[] 1 items
0
- nodata 0
- data_type "uint16"
- bits_per_sample 15
- spatial_resolution 20
- scale 0.0001
- offset 0
roles[] 2 items
- 0 "data"
- 1 "reflectance"
visual-jp2
- href "s3://sentinel-s2-l2a/tiles/34/S/EH/2022/1/23/0/TCI.jp2"
- type "image/jp2"
- title "True color image"
eo:bands[] 3 items
0
- name "red"
- common_name "red"
- description "Red (band 4)"
- center_wavelength 0.665
- full_width_half_max 0.038
1
- name "green"
- common_name "green"
- description "Green (band 3)"
- center_wavelength 0.56
- full_width_half_max 0.045
2
- name "blue"
- common_name "blue"
- description "Blue (band 2)"
- center_wavelength 0.49
- full_width_half_max 0.098
proj:shape[] 2 items
- 0 10980
- 1 10980
proj:transform[] 6 items
- 0 10
- 1 0
- 2 499980
- 3 0
- 4 -10
- 5 4300020
roles[] 1 items
- 0 "visual"
wvp-jp2
- href "s3://sentinel-s2-l2a/tiles/34/S/EH/2022/1/23/0/WVP.jp2"
- type "image/jp2"
- title "Water vapour (WVP)"
proj:shape[] 2 items
- 0 5490
- 1 5490
proj:transform[] 6 items
- 0 20
- 1 0
- 2 499980
- 3 0
- 4 -20
- 5 4300020
raster:bands[] 1 items
0
- nodata 0
- data_type "uint16"
- bits_per_sample 15
- spatial_resolution 20
- unit "cm"
- scale 0.001
- offset 0
roles[] 2 items
- 0 "data"
- 1 "reflectance"
- collection "sentinel-2-l2a"
Prepare bounding box and create the dataset using StacSTAC#
proj = items[0].properties["proj:code"]
poidf = gpd.GeoDataFrame(
pd.DataFrame(),
crs="EPSG:4326",
geometry=[Point(lon, lat)],
).to_crs(proj)
coords = poidf.iloc[0].geometry.coords[0]
# Create bounds in projection
size = 2047
gsd = 20
bounds = (
coords[0] - (size * gsd) // 2,
coords[1] - (size * gsd) // 2,
coords[0] + (size * gsd) // 2,
coords[1] + (size * gsd) // 2,
)
assets = ["blue", "green", "red", "nir","swir16", "scl"]
ds = stackstac.stack(
items,
assets=assets,
bounds=bounds, # pyright: ignore
resolution=gsd,
epsg=int(proj.split(":")[-1]),
dtype="float64", # pyright: ignore
rescale=False,
snap_bounds=True,
resampling=Resampling.nearest,
chunksize=(1, 1, 512, 512),
)
cloud_values = [0, 1, 2, 3, 8, 9, 10] # cloud, shadows, cirrus, etc.
scl_mask = ds.sel(band="scl")
bands = ds.sel(band=["blue", "green", "red", "nir","swir16"])
valid_mask = ~scl_mask.isin(cloud_values)
bands_masked = bands.where(valid_mask)
median_ds = bands_masked.groupby("time.month").median("time", skipna=True)
median_ds = median_ds.fillna(0)
median_ds
<xarray.DataArray 'stackstac-432af8210ba1d087a72b9ed0b155cf6d' (month: 1,
band: 5,
y: 2048, x: 2048)> Size: 168MB
dask.array<where, shape=(1, 5, 2048, 2048), dtype=float64, chunksize=(1, 2, 1024, 1024), chunktype=numpy.ndarray>
Coordinates: (12/20)
* band (band) <U6 120B 'blue' ... 'swir16'
* x (x) float64 16kB 5.014e+05 ... 5...
* y (y) float64 16kB 4.254e+06 ... 4...
s2:datatake_type <U8 32B 'INS-NOBS'
earthsearch:boa_offset_applied bool 1B False
s2:processing_baseline <U5 20B '03.01'
... ...
s2:degraded_msi_data_percentage int64 8B 0
processing:software object 8B {'sentinel2-to-stac': ...
mgrs:latitude_band <U1 4B 'S'
s2:product_type <U7 28B 'S2MSI2A'
epsg int64 8B 32634
* month (month) int64 8B 1
Attributes:
spec: RasterSpec(epsg=32634, bounds=(501400, 4213100, 542360, 4254...
crs: epsg:32634
transform: | 20.00, 0.00, 501400.00|\n| 0.00,-20.00, 4254060.00|\n| 0.0...
resolution: 20median_ds = median_ds.compute()
/Users/syam/virtualenvs/myvenv/lib/python3.13/site-packages/dask/_task_spec.py:744: RuntimeWarning: All-NaN slice encountered
return self.func(*new_argspec, **kwargs)
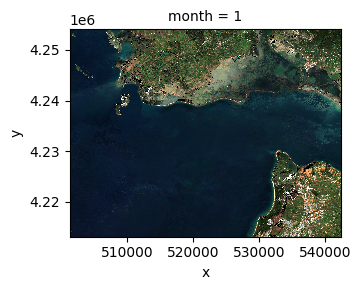
Visualize the ROI#
median_ds.sel(band=["red", "green", "blue"]).plot.imshow(
row="month", rgb="band", vmin=0, vmax=2000, col_wrap=6
)
<xarray.plot.facetgrid.FacetGrid at 0x168ffe510>
Get data as tensor#
pixels = torch.from_numpy(median_ds.data.astype(np.float32))
pixels.shape
torch.Size([1, 5, 2048, 2048])
Convert the tensor to patches of size 5x64x64#
patch_size = 64
stride = 64
batch_size, bands, height, width = pixels.shape
patches = F.unfold(
pixels, kernel_size=patch_size, stride=stride
) # (BATCH, BANDS*PATCH_SIZE*PATCH_SIZE, NUM_PATCHES)
patches = patches.permute(0, 2, 1) # (BATCH, NUM_PATCHES, BANDS*PATCH_SIZE*PATCH_SIZE)
patches = patches.view(
batch_size, -1, bands, patch_size, patch_size
) # (BATCH, NUM_PATCHES, BANDS, PATCH_SIZE, PATCH_SIZE)
patches.shape
torch.Size([1, 1024, 5, 64, 64])
plt.imshow(patches[0,50,0:3,:,:].permute(1,2,0).numpy()/10000*2.5)
plt.axis("off")
plt.show()

patches = patches.reshape(-1, 5, 64, 64)
print(patches.shape)
torch.Size([1024, 5, 64, 64])
Initialize the model#
model = UNet(
in_channels=5,
out_classes=4,
dropout=0.0
)
model_path = "/Users/syam/Documents/code/meditwin/meditwin-summer-school/docs/chapter4/esa_worldcover_classification/models/best_model.pth"
model.load_state_dict(torch.load(model_path,weights_only=True))
model.to("mps")
model.eval()
UNet(
(enc1): Sequential(
(0): Conv2d(5, 64, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1))
(1): BatchNorm2d(64, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(2): ReLU(inplace=True)
(3): Conv2d(64, 64, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1))
(4): BatchNorm2d(64, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(5): ReLU(inplace=True)
(6): Dropout2d(p=0.0, inplace=False)
)
(enc2): Sequential(
(0): Conv2d(64, 128, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1))
(1): BatchNorm2d(128, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(2): ReLU(inplace=True)
(3): Conv2d(128, 128, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1))
(4): BatchNorm2d(128, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(5): ReLU(inplace=True)
(6): Dropout2d(p=0.0, inplace=False)
)
(enc3): Sequential(
(0): Conv2d(128, 256, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1))
(1): BatchNorm2d(256, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(2): ReLU(inplace=True)
(3): Conv2d(256, 256, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1))
(4): BatchNorm2d(256, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(5): ReLU(inplace=True)
(6): Dropout2d(p=0.0, inplace=False)
)
(pool): MaxPool2d(kernel_size=2, stride=2, padding=0, dilation=1, ceil_mode=False)
(bottleneck): Sequential(
(0): Conv2d(256, 512, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1))
(1): BatchNorm2d(512, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(2): ReLU(inplace=True)
(3): Conv2d(512, 512, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1))
(4): BatchNorm2d(512, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(5): ReLU(inplace=True)
(6): Dropout2d(p=0.0, inplace=False)
)
(dec3): Sequential(
(0): Conv2d(768, 256, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1))
(1): BatchNorm2d(256, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(2): ReLU(inplace=True)
(3): Conv2d(256, 256, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1))
(4): BatchNorm2d(256, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(5): ReLU(inplace=True)
(6): Dropout2d(p=0.0, inplace=False)
)
(dec2): Sequential(
(0): Conv2d(384, 128, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1))
(1): BatchNorm2d(128, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(2): ReLU(inplace=True)
(3): Conv2d(128, 128, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1))
(4): BatchNorm2d(128, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(5): ReLU(inplace=True)
(6): Dropout2d(p=0.0, inplace=False)
)
(dec1): Sequential(
(0): Conv2d(192, 64, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1))
(1): BatchNorm2d(64, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(2): ReLU(inplace=True)
(3): Conv2d(64, 64, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1))
(4): BatchNorm2d(64, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(5): ReLU(inplace=True)
(6): Dropout2d(p=0.0, inplace=False)
)
(dec0): Sequential(
(0): Conv2d(64, 32, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1))
(1): BatchNorm2d(32, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(2): ReLU(inplace=True)
(3): Conv2d(32, 32, kernel_size=(3, 3), stride=(1, 1), padding=(1, 1))
(4): BatchNorm2d(32, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True)
(5): ReLU(inplace=True)
(6): Dropout2d(p=0.0, inplace=False)
)
(final): Conv2d(32, 4, kernel_size=(1, 1), stride=(1, 1))
)
Perform inference on the patches#
norm_patches = patches / 10000.0
norm_patches = norm_patches.clip(min=0.0, max=1.0)
norm_patches = norm_patches.to("mps")
with torch.no_grad():
predictions = model(norm_patches)
masks = torch.argmax(predictions, dim=1)
# import gc
# norm_patches = patches / 10000.0
# norm_patches = norm_patches.clip(min=0.0, max=1.0)
# #norm_patches = norm_patches.to("cuda")
# batch_size = 1 # Reduce this to fit memory
# predictions = []
# torch.cuda.empty_cache()
# with torch.no_grad():
# for i in range(0, norm_patches.size(0), batch_size):
# batch = norm_patches[i:i+batch_size].to('cuda')
# out = model(batch)
# predictions.append(out.cpu()) # Move to CPU to save GPU memory
# del batch, out # Free up GPU memory
# gc.collect()
# torch.cuda.empty_cache()
# #with torch.no_grad():
# # predictions = model(norm_patches)
# predictions = torch.cat(predictions, dim=0)
# masks = torch.argmax(predictions, dim=1)
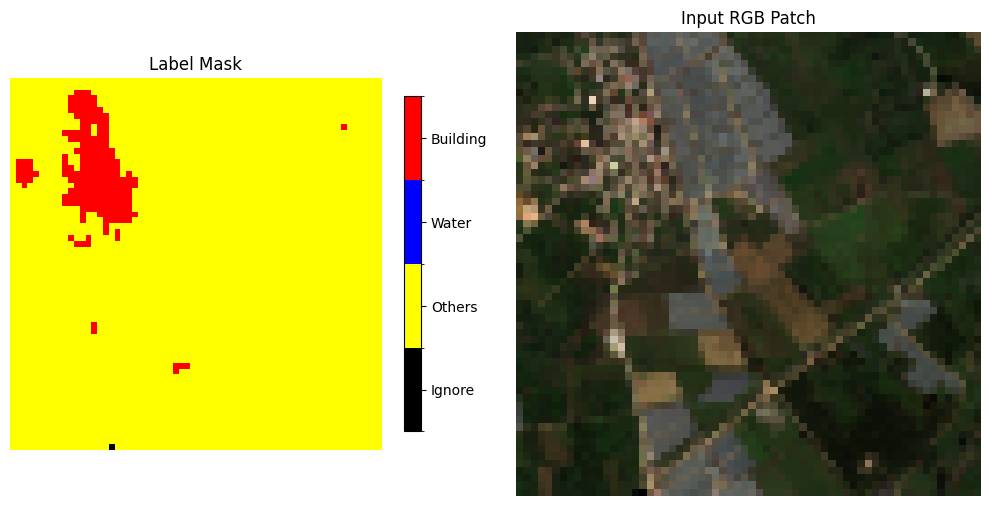
Show an example of the inference result#
idx = 1020 #1000#170#230
cmap = ListedColormap(
[
"black", # 0 = ignore
"yellow", # 1 = other
"blue", # 2 = water
"red", # 3 = building
]
)
bounds = np.arange(0, 5)
norm = BoundaryNorm(bounds, cmap.N)
fig, axs = plt.subplots(1, 2, figsize=(10, 7))
rgb = norm_patches[idx, 0:3].detach().cpu().permute(1, 2, 0).numpy() * 2.5
rgb = rgb[..., [2, 1, 0]]
mask_nodata = rgb[..., 0] == 0
msk = masks[idx].detach().cpu().numpy()
msk[mask_nodata] = 0
msk = msk.astype(np.uint8)
im0 = axs[0].imshow(msk, cmap=cmap, norm=norm)
axs[0].set_title("Label Mask")
cbar = fig.colorbar(im0, ax=axs[0], shrink=0.5, ticks=np.arange(0.5, 4.5))
cbar.ax.set_yticklabels(["Ignore", "Others", "Water", "Building"])
axs[1].imshow(rgb)
axs[1].set_title("Input RGB Patch")
for ax in axs:
ax.axis("off")
plt.tight_layout()
plt.show()
Reconstruct the patch predictions to the original image size#
B, C, H, W = pixels.shape
num_patches_per_img = (H // patch_size) * (W // patch_size)
masks = masks.to(torch.float32)
masks = masks.reshape(B, num_patches_per_img, patch_size, patch_size)
masks = masks.reshape(B, num_patches_per_img, -1).permute(0, 2, 1)
reconstructed = F.fold(
masks, output_size=(H, W), kernel_size=patch_size, stride=stride
)
/Users/syam/virtualenvs/myvenv/lib/python3.13/site-packages/torch/nn/functional.py:5561: UserWarning: The operator 'aten::col2im' is not currently supported on the MPS backend and will fall back to run on the CPU. This may have performance implications. (Triggered internally at /Users/runner/work/pytorch/pytorch/pytorch/aten/src/ATen/mps/MPSFallback.mm:14.)
return torch._C._nn.col2im(
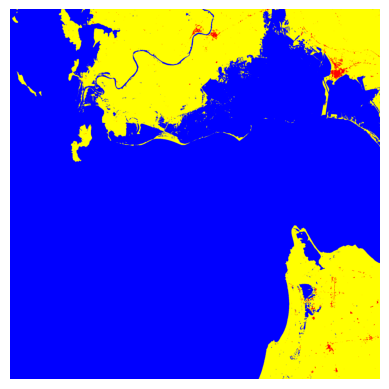
Visualize the full image prediction#
reconstructed = reconstructed.squeeze(1)
reconstructed = reconstructed.cpu().numpy()[0,...]
plt.imshow(reconstructed, cmap=cmap, norm=norm)
plt.axis("off")
plt.show()
Save the prediction as a GeoTIFF#
transform = median_ds.attrs["transform"]
crs = median_ds.attrs["crs"]
with rasterio.open(
"../data/esa_classification_2022_4_classes.tif",
"w",
driver="GTiff",
height=reconstructed.shape[0],
width=reconstructed.shape[1],
count=1,
dtype=reconstructed.dtype,
crs=crs,
transform=transform,
) as dst:
dst.write(reconstructed[np.newaxis, ...])
